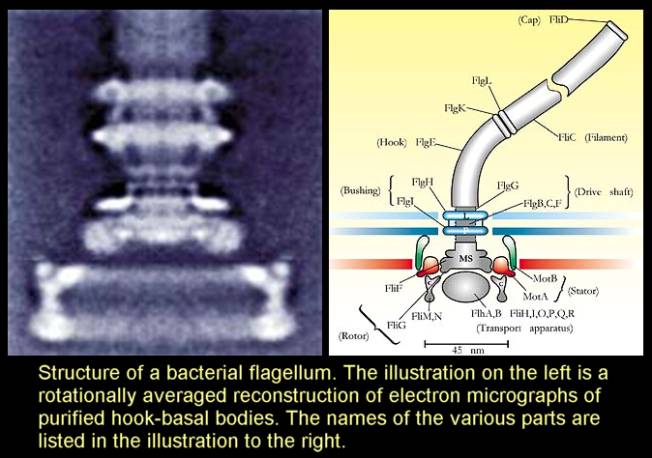
No, they represent potential contributing factors in the space of potential solutions to the circumstantial challenges of survival and reproduction. They’re not targets. They are contingent outcomes of extremely long lineages of blindly samling a huge space of different possible solutions to the problem of survival and reproduction.
A minimum set required to create a molecular machine or other structure would be a more specific target. These targets might never be realized, or they might be found in some organism at some time in history. The challenge is that finding even one protein of a size comparable to a larger flagellar protein would require for 1E132 distinct protein possibilities to exist.
It’s not clear to me why that would be a problem. Can you explain how, if that number is actually required, that would be a problem? Are you saying that, if the average flagellum protein is (say) 300 amino acids long, in order for such a protein to evolve, there would have to be 10^132 functional proteins in that space?
In addition, for evolution to serendipitously stumble across complex molecular machines, the search space would have to be filled with functional possibilities.
Well, yes, and every line of evidence corroborates that being the case. Take a look at the Long-Term Evolution-Experiment with E coli. Pick one of the lineages that has been evolving about 65.000 generations. IIRC something like 600 mutations have been fixed in one of the twelve lineages running in parallel. No two of them are the same today, even though they effectively started from the same clonal population.
Among them are every type of mutation. Duplications, deletions, insertions, transpositions, substitutions. Many of these mutations have happened in all sorts of genetic loci. In regulatory regions, in protein coding genes and what have you. They have happened in genes that affect regulation of cell division, in regions that affect DNA repair pathways, in transporter genes, in metabolic enzymes and so on. There is, evidently, an incredibly vast number of possible ways to further adapt the E coli cells to the flask environment and solve that problem, as each lineage has evolved along some unique mutational trajectory. Some of them have even mutated the same genes, but with different mutations yielding similar fitness gains.
The “problem” of being alive, metabolizing, dividing and reproducing for an E coli cell is a problem with an incredibly complex “solution”. And yet there keeps being new and different ways of contributing to solving that problem while increasing fitness, by genomic changes affecting basically every aspect of E coli molecular physiology.
In other words, vast numbers of other possible configurations of proteins corresponding to locomotive machines must be possible for any to be found. However, if so many possibilities exist, the chances of finding a solution which matches a human-designed outboard rotary motor so closely is nil.
That’s target thinking again. The chances of find any one of that vast number of solutions is nil, but chances are SOME solution will be found. And you can always sit back and try to calculate that after the fact that some complex solution was found, it was an incredibly rare solution out of that vast space. But all of the solutions will be. I’ve seen faces, humans, animals, plants, and even human designs in clouds. What were the odds that a could would come out with that particular shape that day and I would happen to look at it?
What use is that kind of after-the-fact calculation? What was the probability 25 years and 65.000 generations of E coli ago, that one of those 12 lineages of E coli would evolve those 600 particular mutations and find that particular ensemble of mutational solutions to increasing fitness? Having been given any imaginable list of 600 mutations we would have calculated something that would have looked nothing short of statistically impossible, but yet one of those combinations of 600 mutations did evolve.
What does it then accomplish to sit here now and try to work out how unlikely that combination of 600 mutations was? How how significant you happen to find how it looks to you?
However, if so few solutions exist in both the search space for human engineering and for biological systems that such similar designs appear in both engineering and biology, then the chances of nature stumbling upon one of them is nil, since functional combinations would be so rare. In summary, the fact that the flagellum matches so closely with a known design logic means that it can better be understood as a predefined target corresponding to a rotary motor than a haphazardly fit together collection of proteins which just happened to function as a motor.
That makes no sense at all. Anyone can find some personal or superficial significance in any imaginary outcome of the process. Some proteins look like bananas, others look like spaghetti, some look like fans, some look like sex toys. Was that intended too?
Natural selection cannot help add even a single piece to the motor, such as the filament, since adding the piece requires multiple highly-specialized proteins. Until all appear and are then fully integrated into the assembly process, natural selection would work against adding the filament.
When you call them “highly specialized”, you seem to assume without evidence that there is no possibility of functional crossover for a contingently expressed duplicate. As in they did not contribute to some other, more flagellum-related functoin until they fully crossed over in sequence space from one known homologue to another. How do you know that?
The production of a useless protein wastes resources and would be quickly disabled through mutations.
This is typical black-and-white, absolutist, creationist thinking. You see these things in two extremes between fully functional, or so deleterious as to be almost lethal.
It doesn’t have to be useless. And even if it was, there’s a very large crossover zone of lower and situational expression levels which can largely compensate for that, effectively rendering such a duplication invisible to selection, and slowing down the fixation of potentially inactivating mutations. The situation is simply not as black and white as you describe. There is a rather complex relationship between population size, efficiency of selection, and the fitness cost of gene expression. You just can’t make the kind of generalization you do here.
You might try to argue that each protein (filament, assembly cap, 2 joint proteins) served some other role, but the degree of specialization makes such a possibility difficult to justify. And, even if they did play some specific role in some other context, selection would prevent them from being redirected toward a completely different process.
Uhh, no it wouldn’t. Duplications into a region under control of a different promoter, compensatory epistasis, genetic drift, etc. etc. Did you read those two references I linked above? They deal with exactly a scenario such as you describe and what you’re saying just isn’t true.